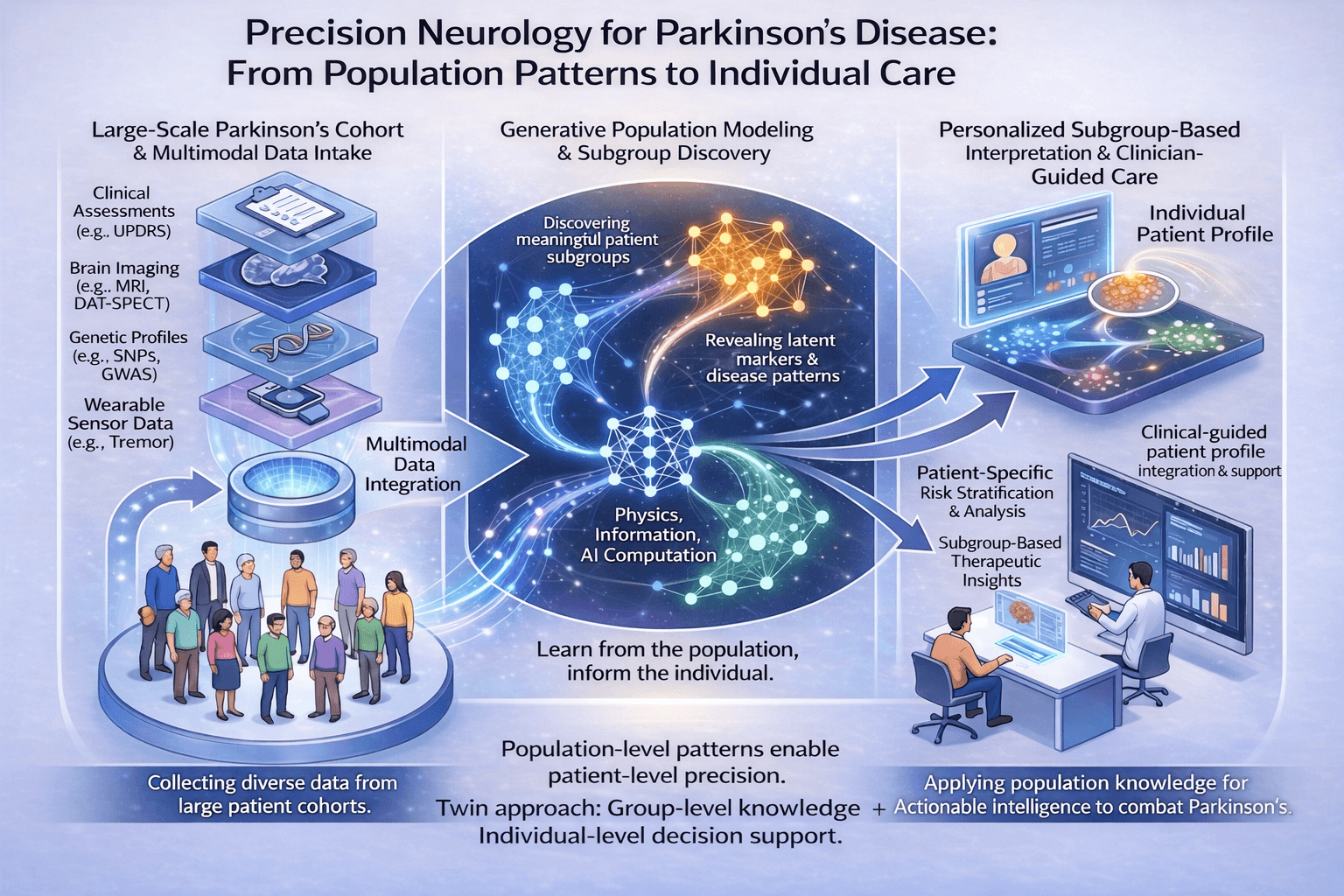
Precision Neurology for Parkinson's Disease
From population patterns to individual care
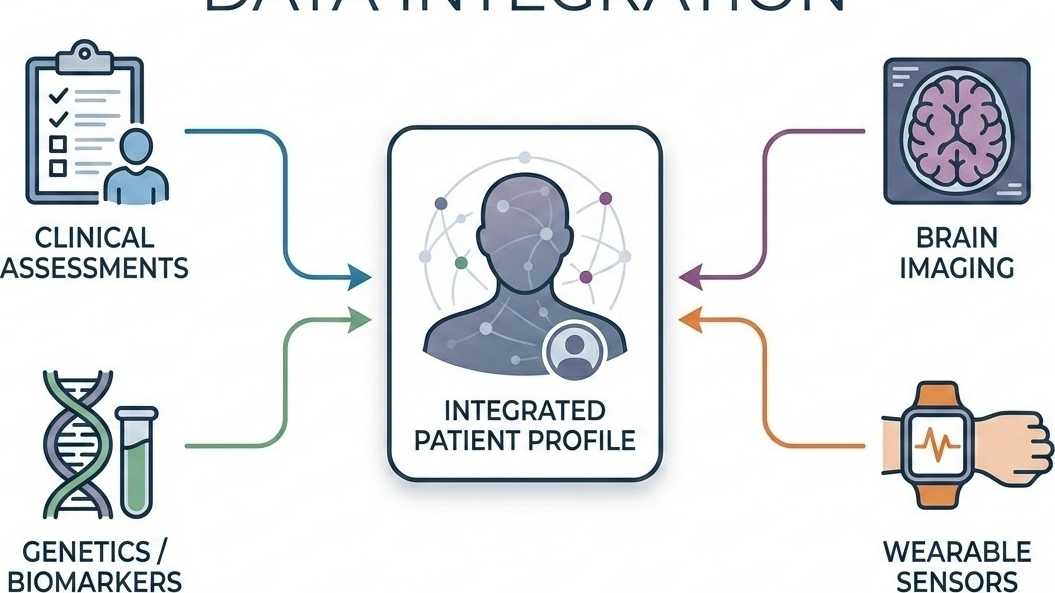
Multimodal Data Integration
We integrate four core data domains — clinical assessments, brain imaging, genetic profiles, and wearable sensor data — from large-scale cohorts to build a comprehensive baseline.
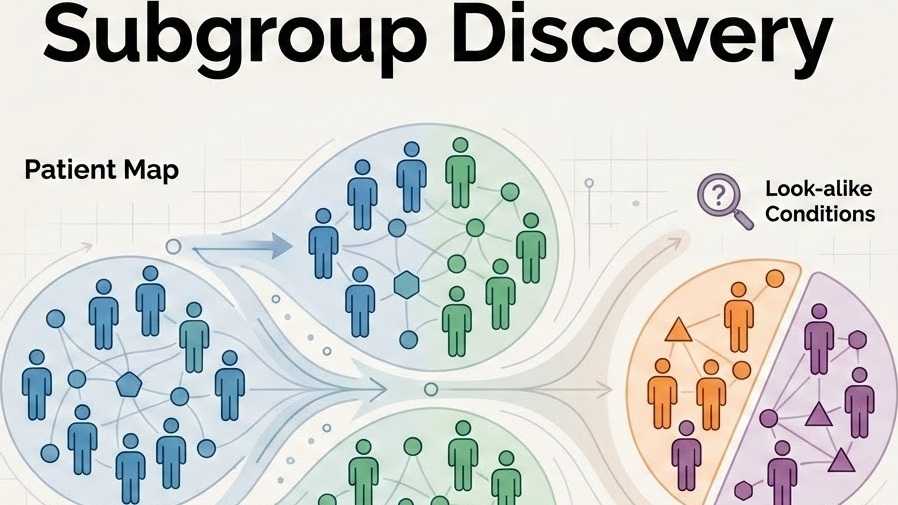
Subgroup Discovery
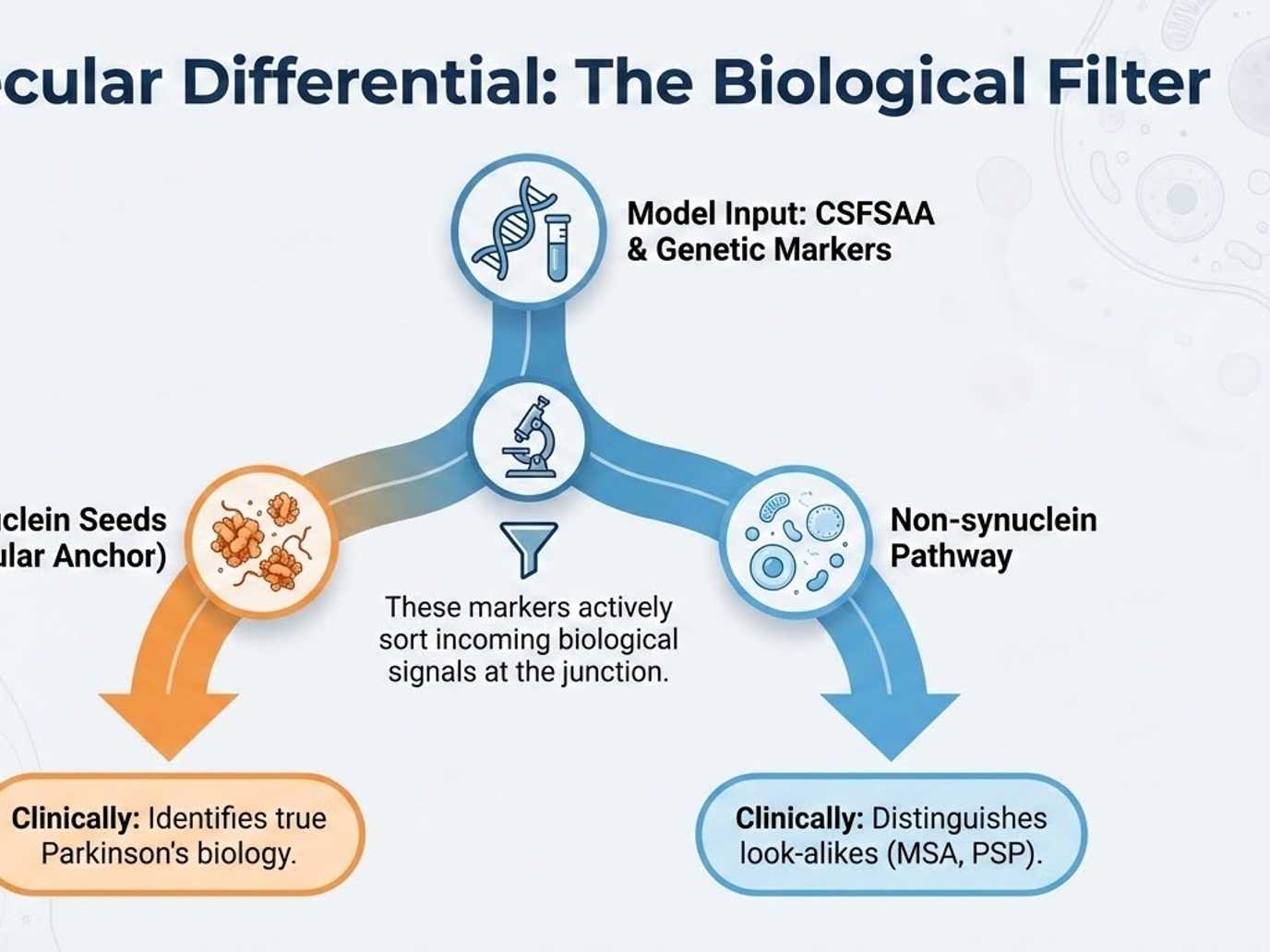
An advanced AI framework analyzes this integrated data to reveal latent markers, differentiating Parkinson's disease from look-alikes and discovering meaningful patient subgroups.
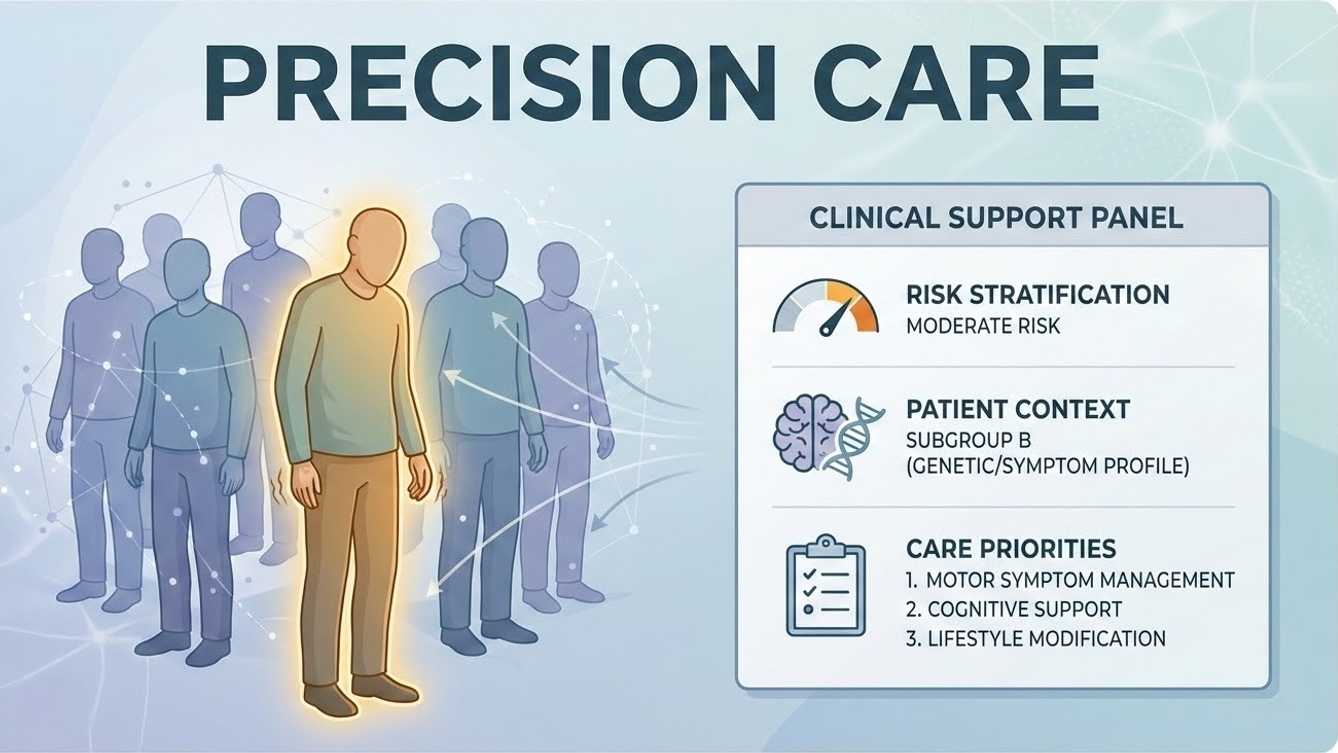
Actionable Precision
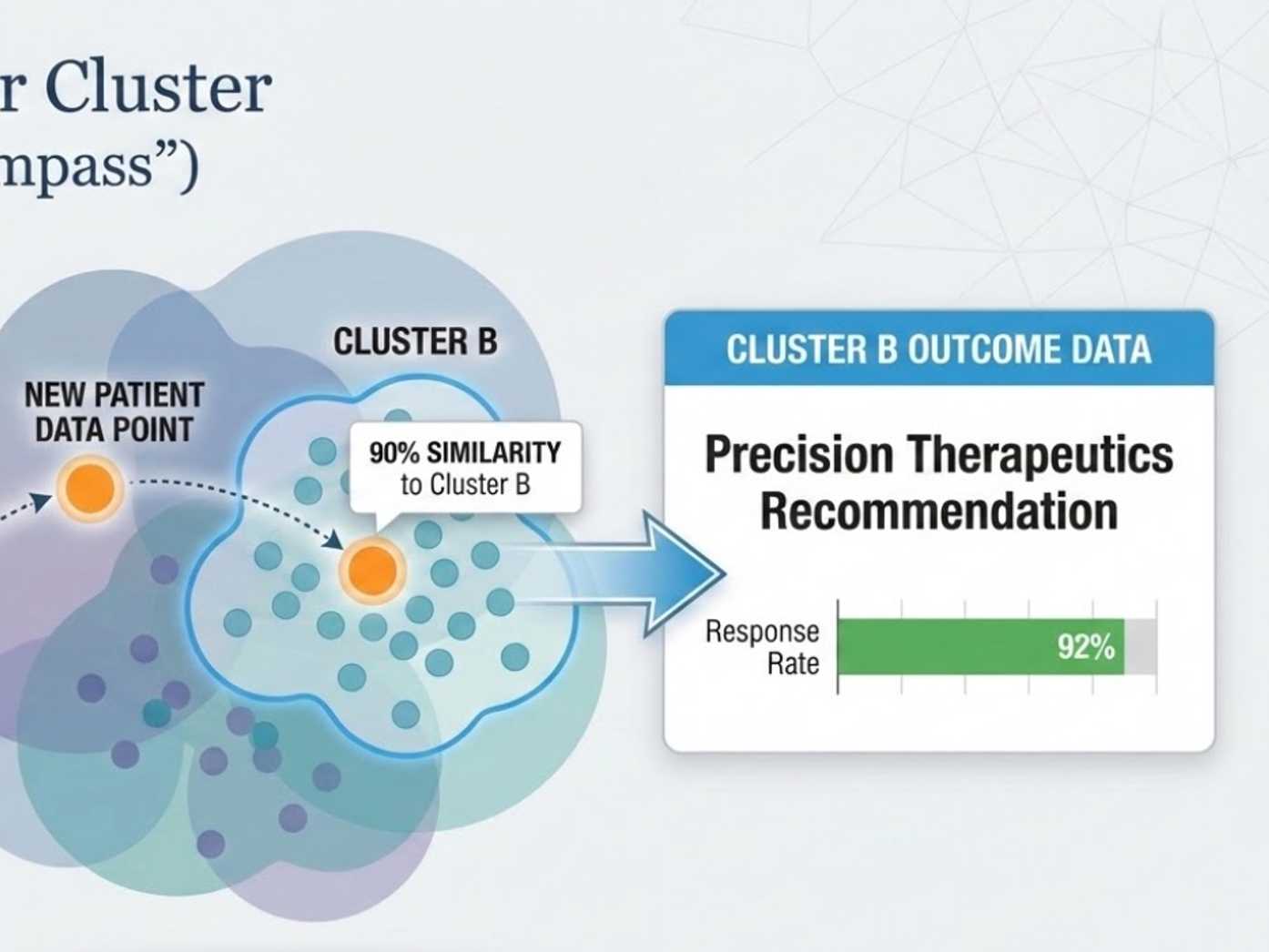
Population-informed models surface patient-specific risk stratifications, subgroup context, and targeted therapeutic priorities.
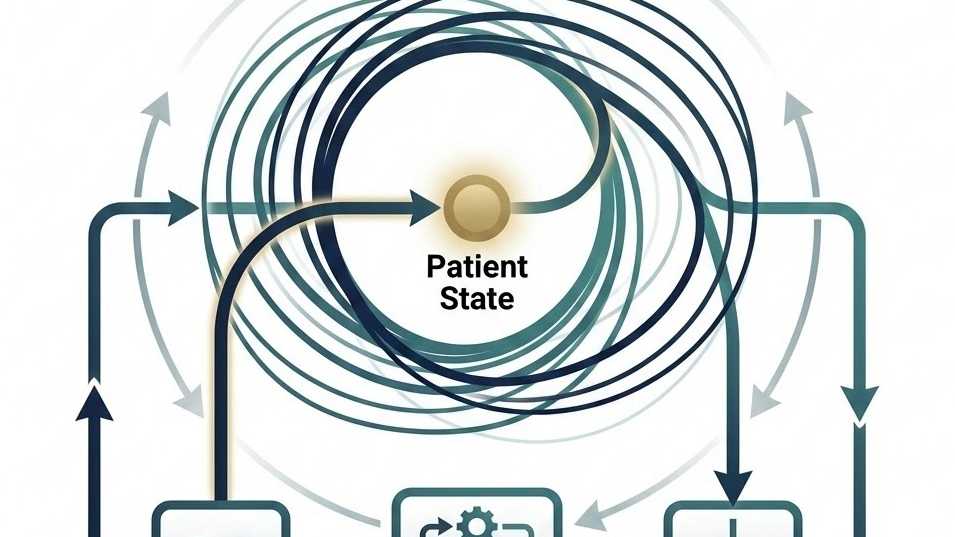
Port Hamiltonian Modeling
We solve the multimodal dynamics with a port-Hamiltonian formulation that preserves structure while supporting stable, interpretable patient-state inference.
What we differentiate
![Differential diagnosis thumbnail]() Differential diagnosis
Differential diagnosisSeed amplification biomarkers help separate Parkinson's disease from look-alikes such as MSA, PSP, and DLB.
![Precision stratification thumbnail]() Precision stratification
Precision stratificationPathway-weighted profiles group patients into biologically distinct Parkinson's subtypes with different trajectories.
![Optimal intervention and decision support thumbnail]() Optimal Intervention and Decision Support
Optimal Intervention and Decision SupportPopulation-informed models translate subgroup context, biomarker burden, and comparable trajectories into intervention priorities and clinician-facing decision support.
Evidence highlights
- Circuit-level imaging signatures reveal patient subtypes with distinct motor and cognitive profiles across six neuroanatomical pathways. View Paper
- Wearable arm-swing asymmetry and dual-task gait cost detect motor-cognitive network involvement non-invasively. View Paper
- LRRK2 G2019S carriers show measurably higher motor severity, enabling genetic risk stratification and personalized counseling. View Paper
- Bayesian clustering identifies four motor phenotypes with transparent model selection via Evidence Lower Bound. View Paper
Sponsors and Funding Partners
Strategic philanthropy, industry collaborations, and clinical trial partnerships accelerate translation of multimodal biomarkers into care. Visit the partners page to learn how funding and clinical collaborators engage, or connect through the resources portal.